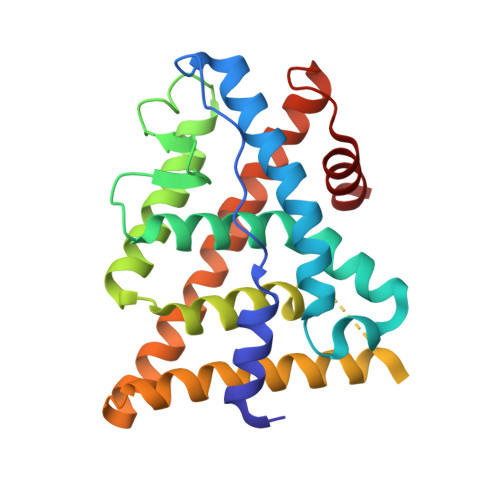
Estrogen receptor ligands: design and synthesis of new 2-arylindene-1-ones
McDevitt, R.E., Malamas, M.S., Manas, E.S., Unwalla, R.J., Xu, Z.B., Miller, C.P., Harris, H.A.(2005) Bioorg Med Chem Lett 15: 3137-3142
- PubMed: 15876535 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2005.04.013
- Primary Citation Related Structures:
1ZAF - PubMed Abstract:
The syntheses of a series of 2-arylindene-1-ones as potent ligands of ERbeta and ERalpha are described. Several compounds exhibited high potency and moderate selectivity for the ERbeta receptor. X-ray and modeling studies were used to understand ligand binding orientation and observed affinity.
- Department of Chemical and Screening Sciences, Wyeth Research, CN 8000, Princeton, NJ 08543-8000, USA. mcdevib@wyeth.com
Organizational Affiliation: